MonaLisa 5.100
Visualization and analysis of functional modules in biochemical networks
Visualization and analysis of functional modules in biochemical networks
Software Specs
Publisher:............ Jens Einloft
License:............... Freeware
File size:.............. 17408 MB
Downloads:.........
Release date:...... 24 Apr 2013
Last update:........ 06 May 2016
Publisher review for MonaLisa 5.100:
Review by: Jens Einloft
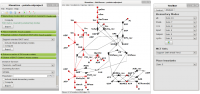
MonaLisa is a Petri net based tool for the analysis of biochemical networks. It comprises an editor supported by an intuitive visualization and animation, and various analysis techniques, focusing on network decomposition, knockout analysis, and other. It includes the computation of elementary modes (T-invariants), P-invariants, Maximal common transition sets (MCT-sets), T-clusters, minimal cut sets (MCS), degree distribution, cluster coefficient distribution.
MonaLisa supports interfaces for different systems biology and graph-theoretic formats: SBML, KGML, PNT, APNN, MetaTool, and PNML.
MonaLisa is written in Java using the JUNG library for the network representation and the Apache Batik library for SVG output.
Requirements:
Java
Operating system:
Windows All
MonaLisa screenshots:
MonaLisa download tags:
Petri net representation biology module Petri net biochemistry
Copyright information:
Based on 0 ratings. 0 user reviews.